miRnomics Atlas
Integration of other omics with miRNAs
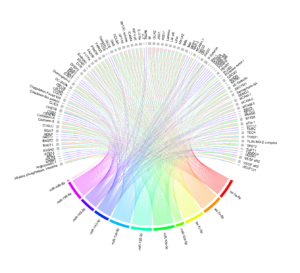
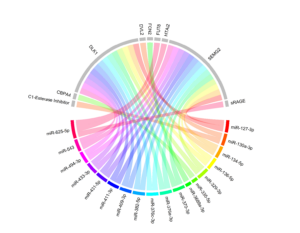
miRNAs have a crucial role in regulating biological processes by modulating the expression of their target genes (mRNAs). Hence, it is of great interest to investigate whether genetic variations linked to expression of miRNAs (miRNA-eQTLs) are associated with the expression of coding genes, including both their host and target genes. To explore this relationship, we conducted an extensive colocalisation analysis to identify overlaps between miRNA-eQTLs and other omics layers, such as gene expression (eQTLs), protein levels (pQTLs), and metabolite levels (mQTLs) in blood/plasma.
For our analysis, we utilized summary statistics from a previous large-scale study (N=31,684) that examined genetic variants influencing mRNA transcript levels, encompassing both cis and trans-eQTLs, in whole blood. We also used GTEx expression data across different tissues (see the reference paper). These statistics enabled us to identify miRNA-eQTLs that impact the expression of other genes. Furthermore, we obtained summary statistics for protein-QTL analyses (pQTLs) from the work by Sun et al. (https://www.phpc.cam.ac.uk/ceu/proteins/) and the SCALLOP consortium (https://zenodo.org/record/2615265/). To identify miRNA-eQTLs affecting metabolic pathways, we leveraged summary statistics from GWAS studies on plasma levels of metabolites. This included over 500 metabolites analyzed in the Metabolon platform by Shin et al. (N=7,824) and nearly 250 metabolites in the Nightingale platform by Kettunen et al. (N>20,000), and data from the UK Biobank study (N=115,078).
DataBase
Search
miRNA
Search all data using a miRNA ID
Protein
Search all data using a protein name
DataBase
Download
Download Description
Here, you can access the summary statistics from our analyses linking miRNA-eQTLs to QTLs in blood/plasma expression of genes, proteins, and metabolites (integrating QTLs data analysis).
Follow US
on These Social Media or Websites